The infographic below gives you the "big picture" on the stupendous hierarchical organization of the human body.

A recent article in MIT News is entitled "Biologist Joey Davis explores how cells build complex structures." We read of a biologist studying how ribosomes get assembled in a cell. At this point the average reader may remember seeing a cell diagram, and the reader may say to himself something like, "Oh, yeah, ribosomes, I remember that there are a few of those little balls in a cell." But the truth is that a typical human cell has millions of ribosomes, and ribosomes are fantastically complex components, like little machines. Ribosomes have the enormously complex function of assembling a huge variety of protein molecules, which are very complex components built from hundreds or thousands of amino acids.
How did the average person get such a wrong idea about ribosomes, the idea that there were only a few of them in the cell, and that they are simple little balls? It is because our biology authorities have done such a poor job of educating us about the vast complexity of cells. Again and again, our biology authorities published misleading diagrams of cells, making it look like cells have only a few parts.
The Google Gemini diagram below discusses some of the great complexities of the work done in the cell by ribosomes.

The diagram has two shortfalls: (1) at the bottom it depicts a cell as enormously less complex than a cell is; (2) at the top it makes ribosomes look vastly less complex than they are. The diagram below helps to show how complex is the structure of ribosomes, which require the special arrangement of very many types of protein molecules:
We see in the diagram above how vastly organized ribosomes are. For a ribosome to be constructed, very many types of proteins must be assembled in just the right way, to produce the "protein factory" that is a ribosome. How does such an assembly occur? Scientists lack any explanation for this miracle of organization. The structure of ribosomes is not specified in DNA or any of its genes. DNA and its genes only contain low-level chemical information, such as which amino acids make up a particular protein. As enormously complex as ribosomes are, their complexity is dwarfed by the complexity of the structures that surround them, the structures called endoplasmic reticulum. A document makes a bungling attempt to explain how the very complex structure of the endoplasmic reticulum arises. It is the vacuous non-explanation of "self organization." The emptiness of the concept is clear from a quote on page 13, where we read, "Self-organization is an interesting concept, but how organelles self-organize is unclear." The last resort of a scientist lacking an explanation for some very high state of organization is to make a vacuous appeal to "self-organization."
Every day that you live, fantastically complex components are magically being assembled in your body, components with a structure that is not specified by DNA or its genes. How this can happen is a mystery a hundred miles over the heads of scientists. They know of no chemistry or physics factors that can explain such assembly.
The wonders seem all the more staggering when we consider the speed at which these miracles of assembly occur. In the MIT News article we get a mention of that. Below is a quote:
"During ribosome assembly, RNA molecules fold themselves into the correct shapes, creating docking sites for proteins to attach. Then, more RNA molecules come in and fold themselves into the structure.
'It’s a beautifully coupled process by which the cell folds hundreds of RNA helices and binds on the order of 50 proteins, and it does it in two minutes from start to finish. E. coli does this 100,000 times per hour, and it’s amazing how rapid and efficient the process is,' Davis says."
How do ribosomes (such fantastically complex components) ever get built? The MIT article offers us no clue, except for two misleading sound bites.
The first misleading sound bite comes in the subtitle of the article, which says this about Joey Davis: "His studies have shed light on the assembly instructions that govern ribosomes, the critical protein-building machines of the cell." That makes it sound as if Davis had studied some assembly instructions for building ribosomes. But no such assembly instructions have ever been discovered, and no such assembly instructions are discussed in the MIT article. DNA and its genes have no assembly instructions for building ribosomes or any other type of organelle.
The second misleading sound bite comes when Davis makes a mention of evolution, without giving any specifics. He says, "It appears that evolution has selected pathways that aren’t strictly ordered in the way we would think about an assembly line, where you always put in one component, then the next, and then the next. " All uses of the phrase "evolution has selected" are misleading, as Darwinian evolution is a mindless process, and only conscious entities can select things. And claims about Darwinian evolution long ago do nothing to explain how ribosomes could get assembled right now in your body. If DNA contained some instructions for how to build ribosomes, then you might be able to make some farfetched appeal to lucky DNA mutations long ago that somehow gave us instructions in DNA for how to build ribosomes. But DNA and its genes do not contain instructions for how to build ribosomes or any other type of organelle in a cell.
Darwinism and claims about evolution are useless in explaining the wonders of biochemistry. That is why Darwin and evolution get virtually no mention in biochemistry textbooks. I documented this reality in my post "The Negligible Presence of Evolutionary Explanations in Six Biochemistry Textbooks," which documents the almost complete lack of mention of evolution, natural selection and Darwin in several long biochemistry textbooks.
Somehow all these marvelously fine-tuned components bigger than protein molecules get assembled in our body, in a way that scientists cannot credibly explain. No assembly instructions for such components and systems have ever been discovered in the body. These miracles of assembly are sometimes very fast and sometimes slow. The progression from a speck-sized zygote to a full human body over the nine months of pregnancy is a miracle of organization, but one that is relatively slow. Conversely, in your body there is constantly occurring very fast miracles of assembly, such as the "two minutes" marvel of assembly mentioned in the quote above.
Some of these miracles of assembly are the construction of protein complexes. Protein complexes are teams of different types of proteins. Proteins somehow assemble into very complex components called protein complexes, which are sometimes so complex they are commonly called "molecular machines" by scientists. Below are some quotes in which scientists confess their lack of understanding of how protein complexes form:
- "The majority of cellular proteins function as subunits in larger protein complexes. However, very little is known about how protein complexes form in vivo." Duncan and Mata, "Widespread Cotranslational Formation of Protein Complexes," 2011.
- "While the occurrence of multiprotein assemblies is ubiquitous, the understanding of pathways that dictate the formation of quaternary structure remains enigmatic." -- Two scientists (link).
- "A general theoretical framework to understand protein complex formation and usage is still lacking." -- Two scientists, 2019 (link).
- "Most proteins associate into multimeric complexes with specific architectures, which often have functional properties like cooperative ligand binding or allosteric regulation. No detailed knowledge is available about how any multimer and its functions arose during historical evolution." -- Ten scientists, 2020 (link).
- "Protein assemblies are at the basis of numerous biological machines by performing actions that none of the individual proteins would be able to do. There are thousands, perhaps millions of different types and states of proteins in a living organism, and the number of possible interactions between them is enormous...The strong synergy within the protein complex makes it irreducible to an incremental process. They are rather to be acknowledged as fine-tuned initial conditions of the constituting protein sequences. These structures are biological examples of nano-engineering that surpass anything human engineers have created. Such systems pose a serious challenge to a Darwinian account of evolution, since irreducibly complex systems have no direct series of selectable intermediates, and in addition, as we saw in Section 4.1, each module (protein) is of low probability by itself." -- Steinar Thorvaldsen and Ola Hössjerm, "Using statistical methods to model the fine-tuning of molecular machines and systems," Journal of Theoretical Biology.
Some such as Hume trying to discredit the idea of miracles have defined a miracle as a violation of the laws of nature. That is not a good definition of "miracle." Here is a good definition of "miracle": a miracle is something (not explained by known laws of physics or chemistry or common human agency) that occurs without visible or known agency, and which would be so improbable to occur by unguided chance that its probability of accidentally occurring is for all practical purposes zero. For example, if you were to take a pack of 52 playing cards, and throw such cards into the air, and all 52 cards became part of a triangular house of cards, that formation so perfect would be something so unlikely to occur that the probability of it occurring is for all practical purposes zero.
Under this reasonable definition of "miracle," the assembly of every ribosome is a miracle, and the assembly of every other very complex and enormously organized organelle is a miracle, as is the assembly of every type of protein complex involving the "just right" arrangement of many types of proteins that assemble into "molecular machines" fine-tuned for some biochemical task. There are no known laws of physics or chemistry that explain such wonders of organization. The assembly of such components by a chance combinations of protein molecules has a probability that is negligible. Under the same definition of "miracle," the assembly of protein complexes as complex as the proteasome (described in this post's appendix) must also be called miracles.
Every day within your body there are a million such miracles, very many of which involve a kind of warp-speed assembly in which parts magically assemble very, very quickly into functional components, in a way that is not predicted by anything we know about the laws of chemistry or physics, and that is not predicted by anything we know about what is in DNA and its genes. The continual occurrence of such miracles is required for the continuation of your life. The physical origin of your body (involving the nine-month progression from a speck-sized zygote to the vast organization of a full human body) was one miracle, but a slow, gradual miracle. The continuation of your body over the span of decades requires endless millions of other miracles, in which purposeful cell components magically assemble in a way that is utterly beyond any explanation of physics, chemistry or genetics.
I asked Google Gemini to produce an infographic visual explaining an example of a protein complex that assembles very quickly. It gave me the visual below. The nuclear pore complex (NPC) discussed is a very well-organized protein complex consisting of more than 30 types of proteins, arranged in just the right way to achieve a particular hard-to-achieve functional effect. Apparently this nuclear pore complex gets assembled within five minutes. The bottom of the diagram has a few lines trying to explain the speed of assembly, but it is little more than the thinnest hand-waving.

How there occurs such miracles of purposeful assembly at such stunning speeds is a mystery a thousand miles over the heads of scientists. When materialists claim that all of the processes of life "can be explained in terms of physics and chemistry," they are telling a lie as big as the sky. The continuation of your life requires the daily occurrence of these miracles of purposeful assembly. The progression from a speck-sized zygote to a vastly more organized full human body over nine months is a miracle of organization very far beyond the explanation of biologists. But it is not merely the origin of every human body that is beyond the explanation of materialists: it is also the continued living existence of an adult body that is a miracle beyond their explanation, because of the million microscopic miracles of warp-speed purposeful assembly that must occur every day for a human body to keep living.
Postscript: Today while searching for some more quotes to add to my "Candid Confessions of the Scientists" post (the largest collection anywhere of scientists confessing what they don't know), I found these two quotes. In one, scientists confess they don't understand how mammary glands arise in a developing body; and in the other scientists confess they don't understand how eyes arise in a developing body.
- "A quarter of the way through the twenty-first century, we still lack basic knowledge regarding the formation and function of the organ that gives its name to all mammals, and which provides important health benefits for children and their breastfeeding parent through the creation and delivery of breast milk." -- 3 scientists (link).
- "Despite increasing knowledge of pathways controlling the differentiation of many cell types in the eye, we still lack a basic understanding of the mechanisms controlling its morphogenesis." - 3 scientists (link).
Also I read today a paper making it clear that contrary to boasts in the press, the AlphaFold2 software does not actually solve the protein folding problem, the problem of how protein molecules almost instantly acquire very complicated 3D shapes needed for their function. The year 2026 paper states, "The explanatory scientific understanding of the protein folding problem is thus not directly advanced by AF2 [AlphaFold2]." Later the same paper says, "The protein folding problem remains unsolved." Instead, the AlphaFold2 software makes progress on a different problem, properly described as the protein structure prediction problem, which is the problem of predicting the 3D shape of a protein molecule from its amino acid sequence. Whenever one of the more complex types of protein molecules almost instantly takes the very complex 3D shape needed for its function, that is another example of a miracle of warp-speed purposeful assembly.
Appendix A: The Proteasome Molecular Machine
Below is one example of the many types of protein complexes that seem to require miracles of assembly beyond the explanation of scientists. The wikipedia.org article on proteasomes tells us this:
"Proteasomes are protein complexes which degrade unneeded or damaged proteins by proteolysis, a chemical reaction that breaks peptide bonds...In structure, the proteasome is a cylindrical complex containing a 'core' of four stacked rings forming a central pore. Each ring is composed of seven individual proteins."
A paper on this topic is entitled "Gates, channels, and switches: elements of the proteasome machine." We read this:
"The proteasome has emerged as an intricate machine that has dynamic mechanisms to regulate the timing of its activity, its selection of substrates, and its processivity. The 19-subunit regulatory particle (RP) recognizes ubiquitinated proteins, removes ubiquitin, and injects the target protein into the proteolytic chamber of the core particle (CP) via a narrow channel."
Another paper is entitled "The 26S Proteasome: A Molecular Machine Designed for Controlled Proteolysis." A page on the site of the Theoretical and Computational Group tells us this:
"Recycling of unneeded protein molecules in cells is performed by a molecular machine called 26S proteasome (Figure 1), which cuts these proteins into smaller pieces for reuse as building blocks for new proteins. Proteins that need to be recycled are labeled by tags made of poly-ubiquitin protein chains. The 26S proteasome machine recognizes and binds to these tags, pulls the tagged protein close, then unwinds it, and finally cuts it into pieces. As the cell's recycling machinery, the 26S proteasome is vital for a variety of essential cellular processes, including protein quality control, cell cycle regulation, adaptive immune response, and apoptosis....The 26S proteasome recruits, unfolds, and degrades poly-ubiquitin tagged proteins through a complex interaction clockwork of over 60 known protein subunits that is driven through ATP hydrolysis."
A scientific paper tells us this:
"The 26S proteasome is a multisubunit complex that catalyzes the degradation of ubiquitinated proteins. The proteasome comprises 33 distinct subunits, all of which are essential for its function and structure."
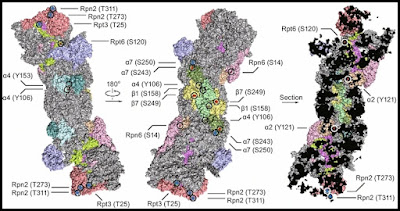
Below is a depiction of the human 26S proteasome structure, one that labels some of its protein parts. We see three different views of the same protein complex, with different protein parts labeled (the Greek letters used stand for alpha and beta parts mentioned in the table below):
Image credit: Xing Guo et. al, link. Below are the number of amino acids involved in these parts, which I looked up using the UniProt online database (you can use the links to check the numbers I have given):
The structure shown above clearly requires several thousands of amino acids that have to be arranged in just the right way. The structure shown above is not specified in DNA, which merely specifies which amino acids make up each of the protein parts. The amino acid information needed to make the structure above (insufficient to specify the total structure) is not at all contiguous in DNA. To assemble the structure above, among other wonders of construction a human body must magically gather genetic information scattered across many different chromosomes in the nucleus, like someone quickly finding just the right 60 loose pages hidden in random books of 46 tall, long bookcases in a library. The table above shows that at least eight of the 23 human chromosome pairs would need to be accessed: Chromosome 1, Chromosome 6, Chromosome 7, Chromosome 11, Chromosome 14, Chromosome 15, Chromosome 17, and Chromosome 20.
Appendix B: The Nuclear Pore Complex
The nuclear pore complex or NPC is a large protein complex found in the "nuclear envelope" that is the outer boundary of the nucleus inside human cells. A science research press release tells us this:
"For structural biologists, the human NPC is a challenging yet exciting 3D puzzle, with around 30 different proteins each present in multiple copies. This amounts to around 1000 puzzle pieces, which form a round core with surrounding flexible parts."
The wikipedia.org article on this complex states that it consists of "456 individual protein molecules, and 34 distinct nucleoporin proteins." So the complex apparently requires 34 types of protein molecules. The article tells us that the "principal function of nuclear pore complexes is to facilitate selective membrane transportation of various molecules across the nuclear envelope." This mean that nuclear pore complexes have the extremely complex job of acting like gatekeepers, letting the right kind of molecules get into the nucleus of the cell, and keeping out the wrong type of molecules. The article tells us that there are typically about 1000 of the nuclear pore complexes in every cell. We read of some impressive functionality of these nuclear pore complexes:
"Notably, the nuclear pore complex (NPC) can actively mediate up to 1000 translocations per complex per second. While smaller molecules can passively diffuse through the pores, larger molecules are often identified by specific signal sequences and are facilitated by nucleoporins to traverse the nuclear envelope."
The article (and also the Google Gemini infographic above) tell us that a nuclear pore complex has a molecular weight of about 110 megadaltons. A dalton is the mass equal to a twelfth of the mass of a carbon atom. A protein complex of 110 megadaltons would have the mass of about 9 million carbon atoms. Apparently the proteins that make up this complex are particularly complex proteins. Below are the exact numbers (we may assume that there are multiple instances of such proteins in a nuclear pore complex).
The molecular machinery shown above clearly requires more than 12,000 amino acids that have to be arranged in just the right way, which amounts to a special arrangement of more than 100,000 atoms. The structure of the molecular machinery described above is not specified in DNA, which merely specifies which amino acids make up each of the protein parts. The amino acid information needed to make the structure above is not at all contiguous in DNA. To assemble the structure above, among other wonders of construction a human body must magically gather genetic information scattered across many different chromosomes in the nucleus, like someone quickly finding just the right 34 loose pages hidden in random books of 46 tall, long bookcases in a library. The table above shows that at least nine of the 23 human chromosome pairs would need to be accessed: Chromosome 1, Chromosome 5, Chromosome 6, Chromosome 7, Chromosome 11, Chromosome 12, Chromosome 16, Chromosome 17 and Chromosome 22.

The nuclear pore complex (credit: Protein Data Bank, link)